Abstract
What is cancer? How does it develop? Doctors and scientists have asked these questions for hundreds of years to understand cancer and find treatments. In this science project, you can investigate these questions too by building a simple model and exploring how environmental and genetic changes affect the development of cancer.Summary

Objective
Build a simple cancer model to investigate how the chances of developing cancer change depending on environmental and genetic factors.
Introduction
Cancer is not new. Neither are humans' efforts to understand what cancer is and find ways to treat and prevent it. The earliest records of cancer date back to descriptions of tumors from an ancient Egyptian textbook on surgery where it was stated "There is no treatment".
Our understanding of cancer and our ability to treat it has evolved a lot since then. We know today that cancer is the unregulated division and growth of cells due to changes in the DNA. Let's break down what that means. Even under normal, healthy conditions, our bodies are in a constant state of growing, dividing, and replacing older cells. We replace approximately 330 billion cells per day! Despite the high number of cell divisions, the process for each division is highly regulated by the cell cycle. The cell cycle provides checks and balances to make sure that cells divide only when they get the right signals to do so and that during division the DNA in the cell is copied and packaged correctly. If mistakes are made during cell division, it is often noticed, and the problematic cells are carefully killed off in a process known as apoptosis. If a mis-divided cell escapes apoptosis, it is often noticed by the immune system and destroyed.
Despite all these checks and balances, things occasionally go wrong and a cell with a mutation (change in its DNA) escapes both apoptosis and the immune system. Everyone has cells in their body with mutations. The older you are, the more mutated cells you are likely to have.
Most mutations do not matter as they do not change how a gene functions. As more cell divisions occur and more mutations accumulate, it may be that (just by chance) some of the mutations do change gene function. If they change genes that control how the cell divides and grows and those changes go unnoticed, the cells may start to grow in an uncontrolled manner—this is cancer.
To understand in detail how diseases like cancer develop and explore treatment options, scientists and doctors create mathematical disease models. There are four steps to mathematical disease modeling:
- Build the model from existing data. That data could be from real-world observations, data from experiments, or a combination of both. The model describes what scientists and doctors think happens with the disease on a biological level.
- Validate the model. The model is tested to see if it makes sense. If the model outputs results that match up with what we observe about the disease in the real world, then the model is considered valid (representative of the truth). If not, the model is revised until it matches the real-world observations.
- Use the model. The model is used to make predictions or run experiments related to the disease, its spread, and/or how to treat it.
- Revise or retire the model. As new observations and data are gathered, the model may need to be revised or retired altogether in favor of a new model.
In this science project, you will build a simple model of how cancer develops using data from published scientific papers and basic arithmetic (addition, subtraction, multiplication, and division). You will use the model to compare how many cell cycles it takes for cancer to develop with and without additional genetic and environmental factors. You will compare the information from your simple model with real-world data to see if it is valid or not.
Terms and Concepts
- Cancer
- Cell cycle
- Apoptosis
- Mutation
- Gene
- Mathematical disease model
- Model validation
Questions
- What is the cell cycle?
- What happens to a cell if it fails to go through the cell cycle properly?
- What do scientists think causes cancer?
Bibliography
Genes, Environment, and Cancer
- HHMI Staff. (n.d.) The Eukaryotic Cell Cycle and Cancer. HHMI Biointeractive. Retrieved September 7, 2023.
- Cancer.Net Editorial Board. (2022, November). Genes and Cancer. American Society of Clinical Oncology. Retrieved September 7, 2023.
Disease Modeling of Cancer
- University of Washington Perspectives. (2021, August 12). Treating Cancer Through Math. Retrieved September 7, 2023.
- Michigan Minds Special Series: Women in STEM. (2021, February 10). Developing Data-Driven Mathematical Models to Study Cancer. University of Michigan Public Engagement & Impact. Retrieved September 7, 2023.
Materials and Equipment
- Computer with an internet connection
- Lab notebook
Experimental Procedure
Building the Starting Cancer Disease Model
- You will be building your cancer disease model using a simple programming language called Scratch. Create a Scratch account.
- Optionally, if you have never used Scratch before, you can follow some of these Scratch programming tutorials to learn how to use it, or look over this Getting Started with Scratch reference guide.
- Figures 1 and 2 show all the things our basic cancer disease model needs to represent.
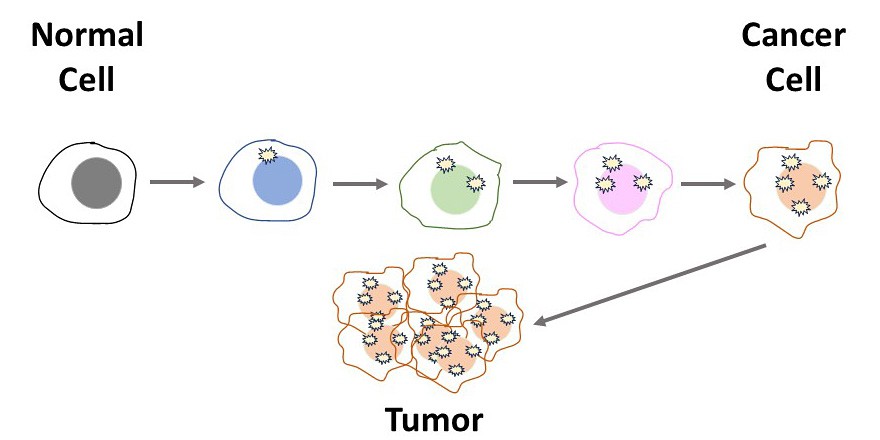
- In Figure 1 we see that when cells divide, the DNA from the original (mother) cell is unchanged but the daughter cell may accumulate mutations. In each cell division, more mutations may accumulate in the daughter cells until finally there are enough mutations that a cancer cell develops. It takes many cell divisions to get a cancer cell.
- In Figure 2 we zoom in to take a closer look at what happens during each cell division.
- Gene mutations arise in the daughter cell. In our cancer model, we will need to assign a normal mutation rate.
- The mutated daughter cell may trigger apoptosis. If it does, it dies. If it does not trigger apoptosis, it continues on. If it continues on, the immune system may detect and destroy the mutated daughter cell. In our cancer model, we can simplify and combine these two events into a single chance of escaping cell death.
- If the mutated daughter cell escapes cell death, it can go through the cell cycle again. If it does, the resulting daughter cell will accumulate even more mutations. In our cancer model, we will need to assign a mutation threshold. If the number of mutations in a daughter cell is greater than the threshold, it will be considered a cancer cell.
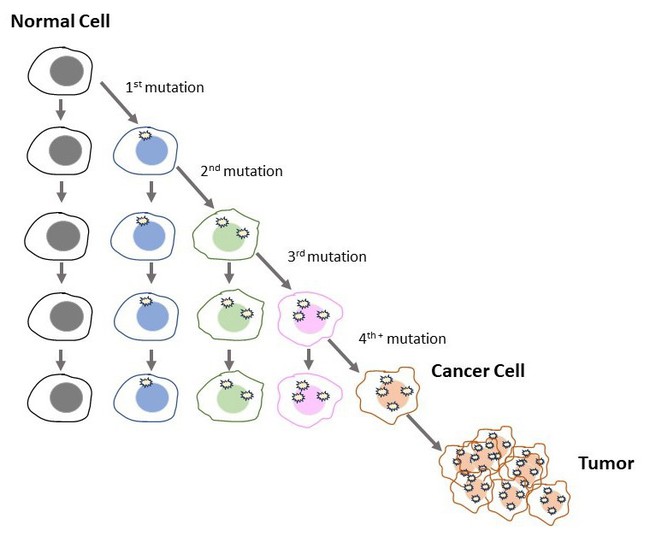 Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Figure 1. Cancer comes from the accumulation of mutations over many cell cycles.
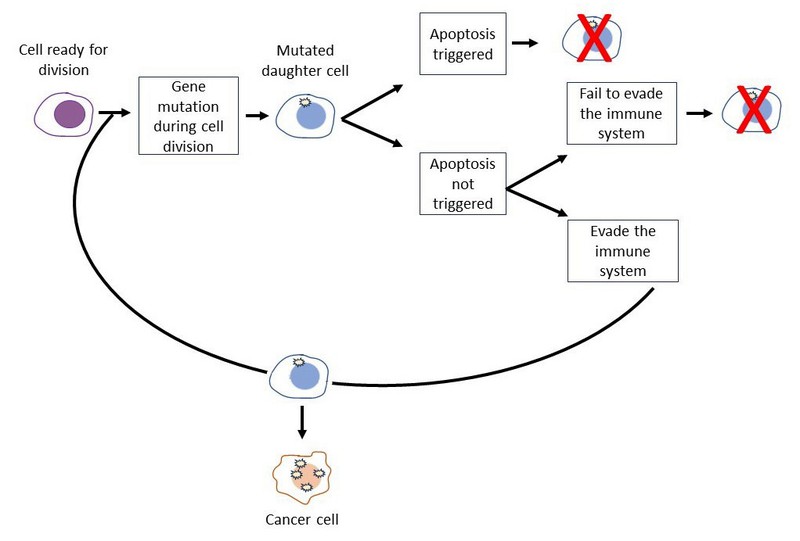 Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Figure 2. This diagram shows how a cell could accumulate mutations through cell division.
- Now that we have outlined what our model does and the data we need to plug in, we can look in the literature to see if that data exists. We have done this step for you in Table 1. If you want to build a more complex model (some suggestions for this can be found in the Variations), you will need to find additional data.
| Description | Data | Reference | Notes |
|---|---|---|---|
| Normal mutation rate for human somatic (not eggs or sperm) cells | 1.2 mutations in the genome per cell division | Lee-Six, Henry et al. Population dynamics of normal human blood inferred from somatic mutations. Nature vol. 561,7724 (2018): 473-478. | |
| Cancer mutation threshold | 1-10 depending on the type of cancer | Martincorena I et al. Universal Patterns of Selection in Cancer and Somatic Tissues. Cell 2017;171;5;1029-1041.e21 | We have chosen to use 10 in the example model we provide. This is a simplification since the research says that mutations in 1-10 specific genes drive each type of cancer. |
| Rate of apoptosis | Not given | N/A | In our example model, we have chosen to combine these and make a guess that mutated daughter cells escape cell death approximately 0.1% of the time. You can adjust this based on your research. |
| Rate of immune cell evasion | Not given | N/A | |
| Mutation rate due to smoking cigarettes | 1 mutation per lung cell per 50 cigarettes | Klein, Alice. November 3, 2016. Every 50 cigarettes smoked cause one DNA mutation per lung cell. New Scientist. | |
| Mutation rate due to an inherited defect in BRCA1 or BRCA2 gene | 7 times higher than the normal mutation rate in breast tissue. | Zámborszky, J. et al. 2017. Loss of BRCA1 or BRCA2 markedly increases the rate of base substitution mutagenesis and has distinct effects on genomic deletions. Oncogene, 36(6), 746-755. |
- You are now ready to build the starting model. Follow steps six through ten to write the starting model yourself, or you can remix this Science Buddies Scratch Project.
- Open Scratch and create five variables:
- Mutation Rate: use this variable to set the rate at which daughter cells accumulate mutations every cell division.
- Cancer Mutation Threshold: use this variable to set the number of mutations a daughter cell must accumulate to become a cancer cell.
- Total Number of Mutations: use this variable to count how many mutations are in the DNA of the cells in your model.
- Number of Cell Cycles: use this variable to count how many cell cycles have passed before a cancer cell appears.
- Cell death: use this to set the chance of a mutated daughter cell escaping cell death.
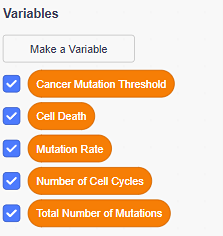 Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Figure 3. Scratch blocks for the five variables needed to build the starting model. - Set the starting values for all your variables. The Mutation Rate and Cancer Mutation Threshold should be set according to Table 1. The rest of the variables should be set to zero.
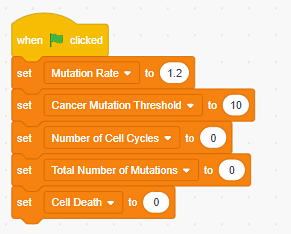 Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Figure 4. Scratch code showing the starting values of the variables. - Program what happens during a single cell division. One possible solution is shown in Figure 5.
- The Number of Cell Cycles increases by one.
- The Total Number of Mutations in the daughter cell increases by the Mutation Rate.
- We check to see if the cell survives Cell Death.
- As explained in Table 1, we have set the chances of escaping Cell Death to 0.1% or 1 in 1000. You can do the same or look for a value based on real-world data in the literature.
- One way to represent a 0.1% chance of escaping cell death is to randomly pick a number between 0.000 and 1.000 for the variable Cell Death. If the value that is randomly picked is greater than 0.001 then the daughter cell dies.
- If the daughter cell dies, then we are left with the mother cell which has fewer Total Number of Mutations. To represent this in the model, we must decrease the Total Number of Mutations by the same amount we added in step 7b.
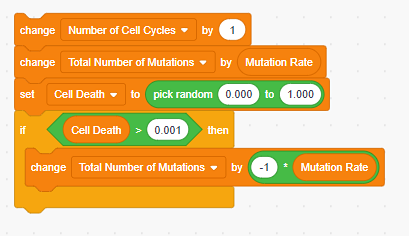 Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Figure 5. Scratch code for what should happen during a single cell cycle. - Now that we have the code for a single cell division, we need to add code to make the cancer model continuously repeat cell cycles until the conditions for cancer are met. In our simple cancer model, we have set a threshold of 10 mutations to be the point at which the daughter cell becomes a cancer cell (see Table 1). One possible solution is shown in Figure 6.
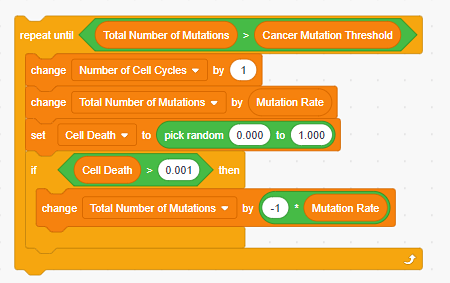 Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Image Credit: Sandra Slutz, Science Buddies / Science Buddies
Figure 6. The scratch code for a single cell cycle will be repeated until the cancer mutation threshold is met. - Optional: You can put in graphics as well to make the program visually appealing and clear to users when the model is finished running.
- Run your model at least ten times. Each time, record in your lab notebook how many cell cycles it takes for a cancer to appear. You can also try running the model here:
Modifying the Cancer Disease Model
- Once you have your starting cancer disease model built, you are ready to modify it to see what happens to the chances of developing cancer under different environmental and genetic conditions.
- Smoking cigarettes is an environmental factor that changes a person's chances of getting lung cancer. Table 1 has information about how smoking affects the mutation rate of cells. Use this information to modify your cancer disease model.
- Run your model for people who smoke at least ten times. Each time, record in your lab notebook how many cell cycles it takes for a cancer to appear.
- Some families with a history of breast cancer have mutations that change the function of the BRCA1 or BRCA2 genes. Using the information in Table 1, modify your starting cancer disease model to see what happens to the chances of getting breast cancer with a BRCA1 or BRCA2 mutation.
- Run your model for people with BRCA1 or BRCA2 mutations at least ten times. Each time, record in your lab notebook how many cell cycles it takes for a cancer to appear.
- Compare your results for the three cancer disease models you built (starting model, smokers' model, and genetic mutation in BRCA1 or BRCA2 model). What is the cell cycle range for each one? What is the average number of cell cycles before cancer appears in each model?
Validating Your Model
- To see if your models are valid (make sense), you will need to compare the results of your models with real-world data. Look up:
- The average age at which smokers versus non-smokers are diagnosed with lung cancer.
- The average age that people with mutations that change the function of BRCA1 or BRCA2 are diagnosed with breast cancer versus people that do not have mutations in either of those genes.
- Do your models match the real-world data? Explain how or how not.
- Optional: Check out the Variations section to see how you can keep improving your models.
Ask an Expert
Global Connections
The United Nations Sustainable Development Goals (UNSDGs) are a blueprint to achieve a better and more sustainable future for all.
Variations
There are many ways to make this model more sophisticated. Try one or more of these suggestions:
- Program your cancer disease model in a different language, like Python, with more capabilities than Scratch. Consider upgrading your model to use a Monte Carlo Simulation. The Monte Carlo Simulation will replace the need to run your model by hand multiple times and will more accurately help you determine the number of cell cycles for each condition (normal, smoking, or BRAC1/2 mutation). To get started, watch this video introduction to the Monte Carlo simulation and consult this python tutorial for Monte Carlo simulations.
- Different cancers have different gene drivers. Model a single type of cancer and the chances of accumulating mutations in the specific genetic drivers for that cancer. This writeup on genetic drivers of cancer is a good place to start your research.
Careers
If you like this project, you might enjoy exploring these related careers:
Related Links
- Science Fair Project Guide
- Other Ideas Like This
- Human Biology & Health Project Ideas
- My Favorites
- Program Your Own COVID-19 or Flu Simulator with Scratch
- Can AI Diagnose Breast Cancer?